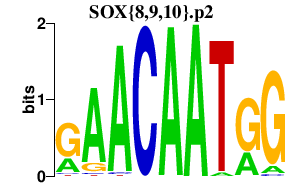
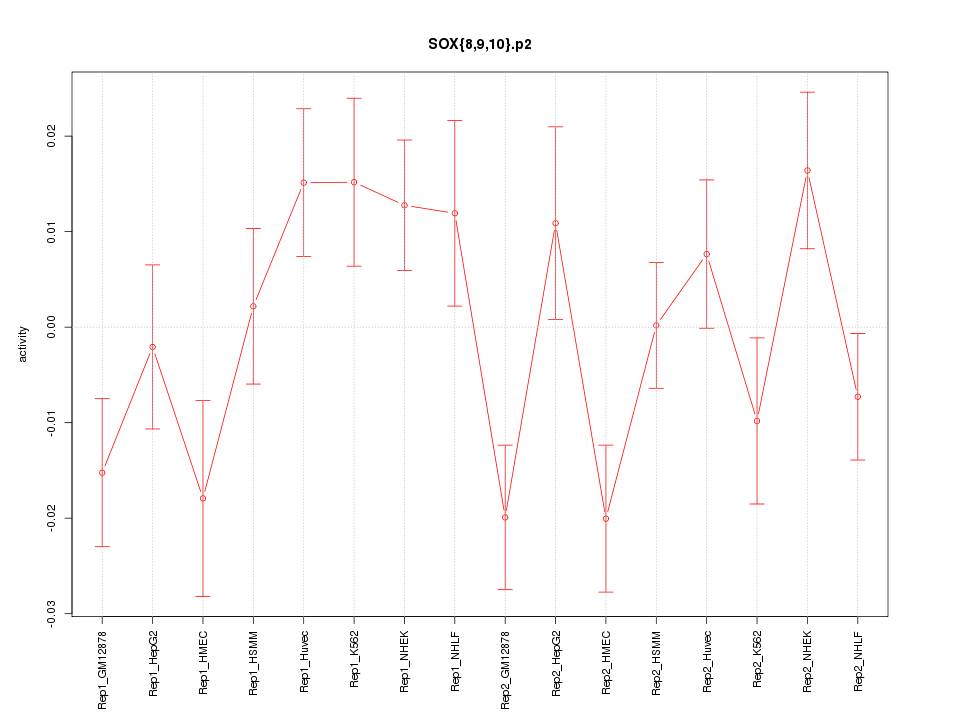
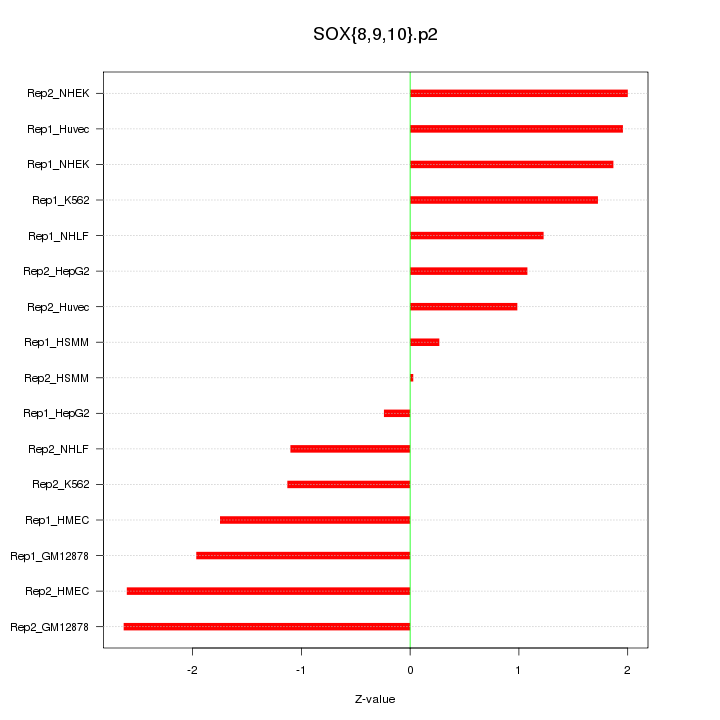
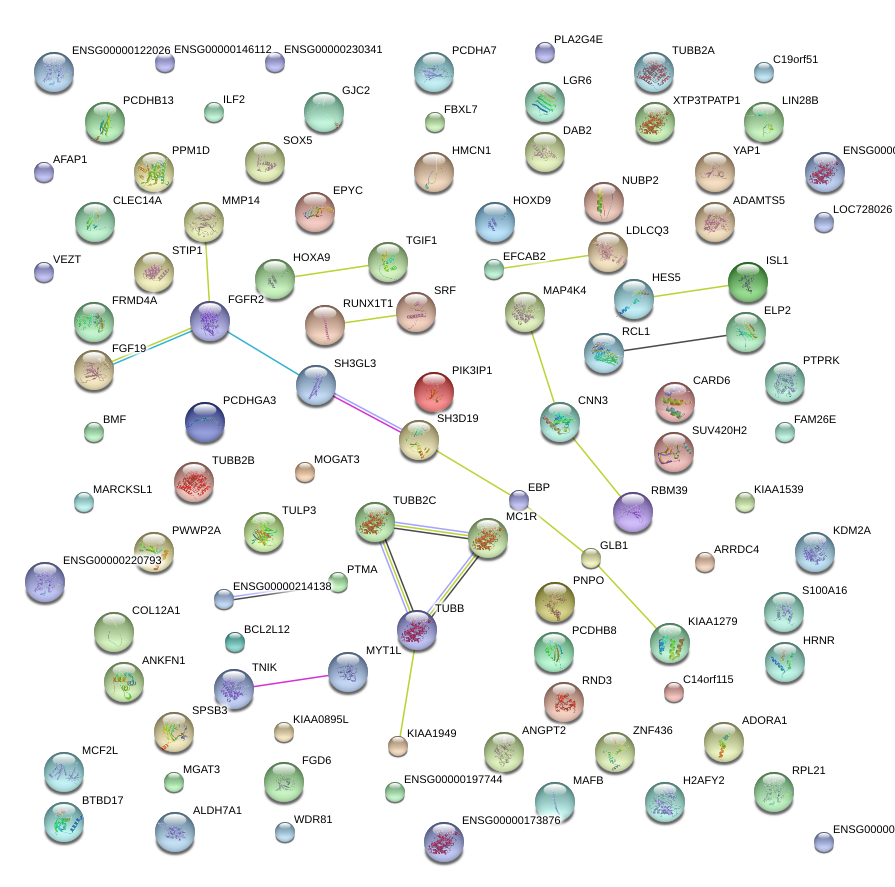
|
chr1_+_183970093
|
1.805
|
NM_031935
|
HMCN1
|
hemicentin 1
|
|
chr6_+_105511615
|
1.541
|
NM_001004317
|
LIN28B
|
lin-28 homolog B (C. elegans)
|
|
chr19_-_60369731
|
1.241
|
NM_178837
|
C19orf51
|
chromosome 19 open reading frame 51
|
|
chr13_+_112704077
|
1.174
|
|
MCF2L
|
MCF.2 cell line derived transforming sequence-like
|
|
chr1_+_200449905
|
1.054
|
NM_001017404
|
LGR6
|
leucine-rich repeat containing G protein-coupled receptor 6
|
|
chr5_-_39461054
|
1.008
|
NM_001343
|
DAB2
|
disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila)
|
|
chr7_-_27171600
|
1.007
|
|
HOXA9
|
homeobox A9
|
|
chr4_-_7991992
|
1.000
|
NM_001134647
NM_198595
|
AFAP1
|
actin filament associated protein 1
|
|
chr11_+_63710206
|
0.995
|
|
STIP1
|
stress-induced-phosphoprotein 1
|
|
chr11_+_63710242
|
0.970
|
|
STIP1
|
stress-induced-phosphoprotein 1
|
|
chr7_-_27171673
|
0.968
|
NM_152739
|
HOXA9
|
homeobox A9
|
|
chr15_-_38188333
|
0.962
|
NM_001003940
|
BMF
|
Bcl2 modifying factor
|
|
chr7_-_27171648
|
0.956
|
|
HOXA9
|
homeobox A9
|
|
chr1_-_151910098
|
0.956
|
NM_004515
|
ILF2
|
interleukin enhancer binding factor 2, 45kDa
|
|
chr1_-_2451543
|
0.954
|
NM_001010926
|
HES5
|
hairy and enhancer of split 5 (Drosophila)
|
|
chr9_+_126460511
|
0.944
|
|
|
|
|
chr14_-_37795290
|
0.936
|
NM_175060
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr11_-_69228286
|
0.933
|
NM_005117
|
FGF19
|
fibroblast growth factor 19
|
|
chr11_+_63710162
|
0.896
|
NM_006819
|
STIP1
|
stress-induced-phosphoprotein 1
|
|
chr14_-_37795001
|
0.893
|
|
CLEC14A
|
C-type lectin domain family 14, member A
|
|
chr9_+_4782833
|
0.889
|
NM_005772
|
RCL1
|
RNA terminal phosphate cyclase-like 1
|
|
chr2_+_232281470
|
0.885
|
NM_001099285
NM_002823
|
PTMA
|
prothymosin, alpha
|
|
chr8_-_93184629
|
0.873
|
NM_001198633
NM_175634
|
RUNX1T1
|
runt-related transcription factor 1; translocated to, 1 (cyclin D-related)
|
|
chr5_+_140772691
|
0.867
|
NM_018913
NM_032090
|
PCDHGA10
|
protocadherin gamma subfamily A, 10
|
|
chr1_-_23568854
|
0.867
|
NM_001077195
|
ZNF436
|
zinc finger protein 436
|
|
chr11_+_66764060
|
0.848
|
|
LOC100131150
|
hypothetical protein LOC100131150
|
|
chr11_+_66764064
|
0.831
|
|
KDM2A
|
lysine (K)-specific demethylase 2A
|
|
chr6_+_30796064
|
0.812
|
|
TUBB
|
tubulin, beta
|
|
chr22_+_38213174
|
0.802
|
NM_001098270
|
MGAT3
|
mannosyl (beta-1,4-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase
|
|
chr5_-_39460787
|
0.791
|
|
DAB2
|
disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila)
|
|
chr5_+_140537554
|
0.787
|
NM_019120
|
PCDHB8
|
protocadherin beta 8
|
|
chr1_+_22107137
|
0.775
|
|
RPL21
|
ribosomal protein L21
|
|
chr21_-_27261276
|
0.763
|
NM_007038
|
ADAMTS5
|
ADAM metallopeptidase with thrombospondin type 1 motif, 5
|
|
chr1_-_32574185
|
0.761
|
NM_023009
|
MARCKSL1
|
MARCKS-like 1
|
|
chr16_-_65774904
|
0.756
|
|
KIAA0895L
|
KIAA0895-like
|
|
chr5_+_140709907
|
0.740
|
NM_018922
NM_032095
|
PCDHGB1
|
protocadherin gamma subfamily B, 1
|
|
chr12_-_94135154
|
0.736
|
NM_018351
|
FGD6
|
FYVE, RhoGEF and PH domain containing 6
|
|
chr19_+_60543032
|
0.734
|
NM_032701
|
SUV420H2
|
suppressor of variegation 4-20 homolog 2 (Drosophila)
|
|
chr16_-_1772305
|
0.727
|
|
SPSB3
|
splA/ryanodine receptor domain and SOCS box containing 3
|
|
chr5_+_40877053
|
0.725
|
NM_032587
|
CARD6
|
caspase recruitment domain family, member 6
|
|
chr8_-_6407882
|
0.717
|
NM_001118887
NM_001118888
NM_001147
|
ANGPT2
|
angiopoietin 2
|
|
chr5_+_50714996
|
0.713
|
|
ISL1
|
ISL LIM homeobox 1
|
|
chr12_-_23995220
|
0.705
|
|
SOX5
|
SRY (sex determining region Y)-box 5
|
|
chr1_-_151910052
|
0.699
|
|
ILF2
|
interleukin enhancer binding factor 2, 45kDa
|
|
chr17_+_56032385
|
0.695
|
|
PPM1D
|
protein phosphatase, Mg2+/Mn2+ dependent, 1D
|
|
chr5_+_74668776
|
0.692
|
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase
|
|
chr10_-_13789628
|
0.692
|
|
FRMD4A
|
FERM domain containing 4A
|
|
chr1_-_151910057
|
0.689
|
|
ILF2
|
interleukin enhancer binding factor 2, 45kDa
|
|
chr5_-_50714836
|
0.687
|
|
LOC642366
|
hypothetical LOC642366
|
|
chr3_-_189354510
|
0.686
|
|
LOC339929
|
hypothetical LOC339929
|
|
chr5_-_39460687
|
0.684
|
|
DAB2
|
disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila)
|
|
chr9_-_35101534
|
0.683
|
|
KIAA1539
|
KIAA1539
|
|
chr17_+_1566566
|
0.682
|
NM_001163673
NM_152348
|
WDR81
|
WD repeat domain 81
|
|
chr12_-_23995109
|
0.675
|
|
SOX5
|
SRY (sex determining region Y)-box 5
|
|
chr14_+_22375753
|
0.669
|
|
MMP14
|
matrix metallopeptidase 14 (membrane-inserted)
|
|
chr5_+_140194152
|
0.666
|
NM_018910
NM_031852
|
PCDHA7
|
protocadherin alpha 7
|
|
chr1_+_226404037
|
0.666
|
NM_020435
|
GJC2
|
gap junction protein, gamma 2, 47kDa
|
|
chr2_+_101680594
|
0.665
|
NM_004834
NM_145686
NM_145687
|
MAP4K4
|
mitogen-activated protein kinase kinase kinase kinase 4
|
|
chr1_-_95165037
|
0.659
|
|
CNN3
|
calponin 3, acidic
|
|
chr5_-_159478967
|
0.657
|
NM_001130864
NM_052927
|
PWWP2A
|
PWWP domain containing 2A
|
|
chr10_+_70418482
|
0.654
|
NM_015634
|
KIAA1279
|
KIAA1279
|
|
chr17_+_43373887
|
0.654
|
NM_018129
|
PNPO
|
pyridoxamine 5'-phosphate oxidase
|
|
chr16_+_1772929
|
0.641
|
NM_012225
|
NUBP2
|
nucleotide binding protein 2 (MinD homolog, E. coli)
|
|
chr6_+_116939500
|
0.636
|
NM_153711
|
FAM26E
|
family with sequence similarity 26, member E
|
|
chr15_+_81907176
|
0.636
|
|
SH3GL3
|
SH3-domain GRB2-like 3
|
|
chr2_-_151052391
|
0.634
|
NM_005168
|
RND3
|
Rho family GTPase 3
|
|
chr22_-_30018329
|
0.634
|
NM_001135911
NM_052880
|
PIK3IP1
|
phosphoinositide-3-kinase interacting protein 1
|
|
chr20_-_33793286
|
0.631
|
|
RBM39
|
RNA binding motif protein 39
|
|
chr6_+_43246897
|
0.630
|
NM_003131
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr5_+_74668748
|
0.629
|
NM_000859
NM_001130996
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase
|
|
chr1_+_201363458
|
0.623
|
NM_000674
NM_001048230
|
ADORA1
|
adenosine A1 receptor
|
|
chr15_+_96304911
|
0.622
|
NM_183376
|
ARRDC4
|
arrestin domain containing 4
|
|
chr17_-_69869552
|
0.621
|
NM_001080466
|
BTBD17
|
BTB (POZ) domain containing 17
|
|
chr12_+_94135645
|
0.617
|
NM_017599
|
VEZT
|
vezatin, adherens junctions transmembrane protein
|
|
chr17_+_51585834
|
0.617
|
NM_153228
|
ANKFN1
|
ankyrin-repeat and fibronectin type III domain containing 1
|
|
chr6_-_128883125
|
0.614
|
NM_002844
NM_001135648
|
PTPRK
|
protein tyrosine phosphatase, receptor type, K
|
|
chr12_-_89922875
|
0.614
|
|
EPYC
|
epiphycan
|
|
chr1_-_151910080
|
0.612
|
|
ILF2
|
interleukin enhancer binding factor 2, 45kDa
|
|
chr6_-_75972473
|
0.609
|
|
COL12A1
|
collagen, type XII, alpha 1
|
|
chr5_+_74668799
|
0.609
|
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase
|
|
chr2_-_2313894
|
0.607
|
NM_015025
|
MYT1L
|
myelin transcription factor 1-like
|
|
chr1_-_151848039
|
0.606
|
|
S100A16
|
S100 calcium binding protein A16
|
|
chr5_-_125958503
|
0.604
|
NM_001182
|
ALDH7A1
|
aldehyde dehydrogenase 7 family, member A1
|
|
chr18_+_3439972
|
0.600
|
NM_173209
NM_173208
NM_003244
|
TGIF1
|
TGFB-induced factor homeobox 1
|
|
chr17_-_76985538
|
0.589
|
|
|
|
|
chr5_+_74668822
|
0.575
|
|
HMGCR
|
3-hydroxy-3-methylglutaryl-CoA reductase
|
|
chr12_-_89922888
|
0.571
|
NM_004950
|
EPYC
|
epiphycan
|
|
chr1_-_151910041
|
0.570
|
|
ILF2
|
interleukin enhancer binding factor 2, 45kDa
|
|
chr10_-_123346148
|
0.565
|
NM_001144915
|
FGFR2
|
fibroblast growth factor receptor 2
|
|
chr6_+_43247168
|
0.563
|
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr12_+_2870255
|
0.562
|
NM_001160408
NM_003324
|
TULP3
|
tubby like protein 3
|
|
chr20_-_38751289
|
0.555
|
NM_005461
|
MAFB
|
v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian)
|
|
chr15_+_96304942
|
0.555
|
|
ARRDC4
|
arrestin domain containing 4
|
|
chr16_-_30841938
|
0.553
|
|
NCRNA00095
|
non-protein coding RNA 95
|
|
chr5_+_15553768
|
0.550
|
|
FBXL7
|
F-box and leucine-rich repeat protein 7
|
|
chr14_+_22375747
|
0.549
|
|
MMP14
|
matrix metallopeptidase 14 (membrane-inserted)
|
|
chr11_+_101488380
|
0.548
|
NM_001195045
|
YAP1
|
Yes-associated protein 1
|
|
chrX_+_48265107
|
0.547
|
NM_006579
|
EBP
|
emopamil binding protein (sterol isomerase)
|
|
chr10_+_71482362
|
0.546
|
NM_018649
|
H2AFY2
|
H2A histone family, member Y2
|
|
chr1_-_23567084
|
0.540
|
NM_030634
|
ZNF436
|
zinc finger protein 436
|
|
chr6_-_30762957
|
0.540
|
|
KIAA1949
|
KIAA1949
|
|
chr11_-_27450808
|
0.537
|
NM_018490
|
LGR4
|
leucine-rich repeat containing G protein-coupled receptor 4
|
|
chr10_+_104253706
|
0.536
|
NM_001178133
NM_016169
|
SUFU
|
suppressor of fused homolog (Drosophila)
|
|
chr20_-_49818306
|
0.535
|
NM_006045
|
ATP9A
|
ATPase, class II, type 9A
|
|
chr6_+_114285209
|
0.532
|
NM_002356
|
MARCKS
|
myristoylated alanine-rich protein kinase C substrate
|
|
chr7_+_16759845
|
0.529
|
NM_014399
|
TSPAN13
|
tetraspanin 13
|
|
chr6_-_75972286
|
0.527
|
NM_004370
NM_080645
|
COL12A1
|
collagen, type XII, alpha 1
|
|
chr11_-_20138445
|
0.526
|
NM_001029865
|
DBX1
|
developing brain homeobox 1
|
|
chr3_+_195336625
|
0.524
|
NM_005524
|
HES1
|
hairy and enhancer of split 1, (Drosophila)
|
|
chr5_-_16989371
|
0.522
|
NM_012334
|
MYO10
|
myosin X
|
|
chr7_-_13997389
|
0.515
|
NM_001163149
|
ETV1
|
ets variant 1
|
|
chr6_-_30763057
|
0.514
|
NM_001134870
|
KIAA1949
|
KIAA1949
|
|
chr9_-_35101592
|
0.514
|
|
KIAA1539
|
KIAA1539
|
|
chr7_-_27206249
|
0.511
|
NM_000522
|
HOXA13
|
homeobox A13
|
|
chr9_+_102275310
|
0.508
|
NM_003692
|
TMEFF1
|
transmembrane protein with EGF-like and two follistatin-like domains 1
|
|
chr18_+_3437607
|
0.508
|
NM_173207
|
TGIF1
|
TGFB-induced factor homeobox 1
|
|
chr7_-_28964186
|
0.507
|
|
|
|
|
chr6_+_17390273
|
0.503
|
NM_001143941
|
RBM24
|
RNA binding motif protein 24
|
|
chr20_-_49818268
|
0.501
|
|
ATP9A
|
ATPase, class II, type 9A
|
|
chr9_+_4782876
|
0.501
|
|
RCL1
|
RNA terminal phosphate cyclase-like 1
|
|
chr2_+_62787534
|
0.499
|
NM_001142614
|
EHBP1
|
EH domain binding protein 1
|
|
chr15_+_76517623
|
0.498
|
|
IREB2
|
iron-responsive element binding protein 2
|
|
chr6_-_42218692
|
0.494
|
NM_001164446
|
C6orf132
|
chromosome 6 open reading frame 132
|
|
chr22_-_27526452
|
0.493
|
|
XBP1
|
X-box binding protein 1
|
|
chr2_-_175337386
|
0.491
|
NM_000079
NM_001039523
|
CHRNA1
|
cholinergic receptor, nicotinic, alpha 1 (muscle)
|
|
chr12_-_89976261
|
0.490
|
NM_007035
|
KERA
|
keratocan
|
|
chr5_+_15553736
|
0.488
|
|
FBXL7
|
F-box and leucine-rich repeat protein 7
|
|
chr7_+_20336849
|
0.486
|
|
ITGB8
|
integrin, beta 8
|
|
chr1_-_226679648
|
0.484
|
NM_003493
|
HIST3H3
|
histone cluster 3, H3
|
|
chr4_+_41309482
|
0.477
|
|
LIMCH1
|
LIM and calponin homology domains 1
|
|
chr4_+_74491272
|
0.477
|
|
ALB
|
albumin
|
|
chr14_+_85066265
|
0.476
|
|
FLRT2
|
fibronectin leucine rich transmembrane protein 2
|
|
chr10_+_104253716
|
0.475
|
|
SUFU
|
suppressor of fused homolog (Drosophila)
|
|
chr7_-_13997574
|
0.473
|
NM_001163148
|
ETV1
|
ets variant 1
|
|
chr11_-_128567302
|
0.473
|
NM_001142685
|
ARHGAP32
|
Rho GTPase activating protein 32
|
|
chr20_-_33793591
|
0.473
|
|
RBM39
|
RNA binding motif protein 39
|
|
chr12_-_24606616
|
0.471
|
NM_152989
|
SOX5
|
SRY (sex determining region Y)-box 5
|
|
chr1_+_15957832
|
0.469
|
NM_017556
|
FBLIM1
|
filamin binding LIM protein 1
|
|
chr3_+_9748769
|
0.467
|
|
BRPF1
|
bromodomain and PHD finger containing, 1
|
|
chr15_+_81906983
|
0.466
|
NM_003027
|
SH3GL3
|
SH3-domain GRB2-like 3
|
|
chr12_+_47498778
|
0.466
|
NM_000725
|
CACNB3
|
calcium channel, voltage-dependent, beta 3 subunit
|
|
chr6_-_64087806
|
0.465
|
NM_001143940
NM_016571
|
LGSN
|
lengsin, lens protein with glutamine synthetase domain
|
|
chr7_+_16652282
|
0.465
|
NM_001159767
NM_014038
|
BZW2
|
basic leucine zipper and W2 domains 2
|
|
chr2_-_217268492
|
0.462
|
NM_000599
|
IGFBP5
|
insulin-like growth factor binding protein 5
|
|
chr17_+_18069593
|
0.460
|
NM_004140
|
LLGL1
|
lethal giant larvae homolog 1 (Drosophila)
|
|
chr11_+_111313067
|
0.460
|
NM_001037954
|
DIXDC1
|
DIX domain containing 1
|
|
chr16_+_81218142
|
0.453
|
|
CDH13
|
cadherin 13, H-cadherin (heart)
|
|
chr20_-_33793593
|
0.453
|
NM_004902
NM_184234
|
RBM39
|
RNA binding motif protein 39
|
|
chr2_+_223244606
|
0.453
|
NM_058165
|
MOGAT1
|
monoacylglycerol O-acyltransferase 1
|
|
chr2_-_119322228
|
0.448
|
NM_001426
|
EN1
|
engrailed homeobox 1
|
|
chr11_-_16380967
|
0.448
|
NM_001145819
NM_017508
|
SOX6
|
SRY (sex determining region Y)-box 6
|
|
chr12_-_11230809
|
0.445
|
NM_181429
|
TAS2R42
|
taste receptor, type 2, member 42
|
|
chr1_-_68470585
|
0.438
|
NM_001002292
NM_001193334
NM_024911
|
WLS
|
wntless homolog (Drosophila)
|
|
chr20_-_33793507
|
0.437
|
|
RBM39
|
RNA binding motif protein 39
|
|
chr17_+_43374021
|
0.437
|
|
PNPO
|
pyridoxamine 5'-phosphate oxidase
|
|
chr18_-_13905534
|
0.436
|
NM_000529
|
MC2R
|
melanocortin 2 receptor (adrenocorticotropic hormone)
|
|
chr11_-_85074848
|
0.435
|
NM_152723
|
CCDC89
|
coiled-coil domain containing 89
|
|
chr5_-_16989290
|
0.433
|
|
MYO10
|
myosin X
|
|
chr19_-_63205948
|
0.433
|
|
ZNF606
|
zinc finger protein 606
|
|
chr6_-_31617704
|
0.426
|
|
DDX39B
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B
|
|
chr6_-_31617748
|
0.425
|
|
DDX39B
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39B
|
|
chr6_+_31190505
|
0.424
|
NM_014068
|
PSORS1C1
|
psoriasis susceptibility 1 candidate 1
|
|
chr17_+_75366524
|
0.423
|
NM_005189
NM_032647
|
CBX2
|
chromobox homolog 2
|
|
chr10_-_75080784
|
0.422
|
NM_024875
|
SYNPO2L
|
synaptopodin 2-like
|
|
chr7_-_5429702
|
0.422
|
NM_001080495
|
TNRC18
|
trinucleotide repeat containing 18
|
|
chr8_-_23619866
|
0.420
|
NM_001136271
|
NKX2-6
|
NK2 transcription factor related, locus 6 (Drosophila)
|
|
chr15_-_62904857
|
0.416
|
NM_025049
|
PIF1
|
PIF1 5'-to-3' DNA helicase homolog (S. cerevisiae)
|
|
chr14_+_38714111
|
0.416
|
NM_002687
|
PNN
|
pinin, desmosome associated protein
|
|
chr15_-_62460622
|
0.415
|
NM_001029989
NM_014736
|
KIAA0101
|
KIAA0101
|
|
chr1_+_23568028
|
0.414
|
|
C1orf213
|
chromosome 1 open reading frame 213
|
|
chr5_+_140719773
|
0.414
|
NM_018923
NM_032096
|
PCDHGB2
|
protocadherin gamma subfamily B, 2
|
|
chr7_-_11838260
|
0.414
|
NM_015204
|
THSD7A
|
thrombospondin, type I, domain containing 7A
|
|
chr6_+_43247179
|
0.413
|
|
SRF
|
serum response factor (c-fos serum response element-binding transcription factor)
|
|
chr6_+_30796123
|
0.413
|
NM_178014
|
TUBB
|
tubulin, beta
|
|
chr18_+_54039779
|
0.413
|
|
NEDD4L
|
neural precursor cell expressed, developmentally down-regulated 4-like
|
|
chr9_+_115303527
|
0.411
|
NM_130795
|
RGS3
|
regulator of G-protein signaling 3
|
|
chr12_+_121885246
|
0.408
|
|
HIP1R
|
huntingtin interacting protein 1 related
|
|
chr12_-_47362254
|
0.407
|
NM_017822
|
C12orf41
|
chromosome 12 open reading frame 41
|
|
chr18_+_18003401
|
0.406
|
NM_005257
|
GATA6
|
GATA binding protein 6
|
|
chr1_-_160260210
|
0.404
|
NM_015441
|
OLFML2B
|
olfactomedin-like 2B
|
|
chr14_+_36196523
|
0.404
|
NM_006194
|
PAX9
|
paired box 9
|
|
chr8_+_121892719
|
0.403
|
|
|
|
|
chr1_-_95165089
|
0.402
|
NM_001839
|
CNN3
|
calponin 3, acidic
|
|
chr11_+_11820096
|
0.399
|
|
USP47
|
ubiquitin specific peptidase 47
|
|
chr4_-_122964428
|
0.398
|
NM_001237
|
CCNA2
|
cyclin A2
|
|
chr11_+_11820041
|
0.395
|
|
|
|
|
chr16_+_30993252
|
0.394
|
|
ZNF646
|
zinc finger protein 646
|
|
chr14_+_62740878
|
0.394
|
NM_020663
|
RHOJ
|
ras homolog gene family, member J
|
|
chr3_-_11598829
|
0.394
|
NM_001128220
|
VGLL4
|
vestigial like 4 (Drosophila)
|
|
chr4_-_159312973
|
0.391
|
|
FAM198B
|
family with sequence similarity 198, member B
|
|
chr5_+_50714714
|
0.390
|
NM_002202
|
ISL1
|
ISL LIM homeobox 1
|
|
chr12_-_106011715
|
0.390
|
NM_004075
|
CRY1
|
cryptochrome 1 (photolyase-like)
|
|
chr6_-_139737477
|
0.389
|
NM_006079
|
CITED2
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2
|
|
chr12_+_11694172
|
0.387
|
|
ETV6
|
ets variant 6
|
|
chr16_+_21518633
|
0.387
|
|
METTL9
|
methyltransferase like 9
|
|
chr5_+_15553539
|
0.386
|
|
FBXL7
|
F-box and leucine-rich repeat protein 7
|
|
chr3_+_85090721
|
0.385
|
NM_001167674
NM_001167675
|
CADM2
|
cell adhesion molecule 2
|
|
chr1_+_159495140
|
0.385
|
NM_001102566
|
PCP4L1
|
Purkinje cell protein 4 like 1
|
|
chr4_-_87500398
|
0.385
|
NM_138980
NM_138982
|
MAPK10
|
mitogen-activated protein kinase 10
|
|
chr16_+_81218074
|
0.384
|
NM_001257
|
CDH13
|
cadherin 13, H-cadherin (heart)
|